|
Layers
Diffusion in heterogeneous environments
|
|
Layers
Diffusion in heterogeneous environments
|
Header file for program fit-layer. More...
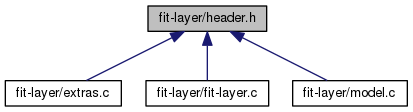
Go to the source code of this file.
Macros | |
| #define | MAXNUM_LINES 10000 |
| Maximum number of lines in input file. | |
| #define | MAX_LINELENGTH 100 |
| Maximum length of any line in input file. | |
| #define | MAXNUM_COMMENTLINES 1000 |
| Maximum number of comment lines of input file to copy to output file. | |
| #define | MAX_COMMAND_LENGTH 1000 |
| Maximum number of characters of command to copy to output file. | |
| #define | FALSE 0 |
| FALSE assigned to 0. | |
| #define | TRUE 1 |
| TRUE assigned to 1. | |
| #define | PI (4*atan(1.0)) |
Sets constant PI to M_PI if that is defined; otherwise, computes PI as 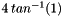 . . | |
| #define | FARADAY 96485.3399 |
| Faraday constant in C/mol. | |
| #define | SMALLNUM 2.2204460492503131e-16 |
| For comparing doubles with 0: Sets SMALLNUM to DBL_EPSILON or to GSL_DBL_EPSILON if they're defined; otherwise set to another small number. | |
| #define | IS_ZERO(x) (fabs(x) < (SMALLNUM) ? (TRUE) : (FALSE)) |
| Evaluates to TRUE if |x| < SMALLNUM, FALSE otherwise. | |
| #define | STREQ(s1, s2) (strcmp(s1,s2) == 0) |
| Compares strings s1 and s2; evaluates to TRUE if strings are equal, FALSE if not. | |
| #define | SQR(x) ((x) * (x)) |
| Computes the square of x. | |
| #define | MAX(x, y) ((x) > (y) ? (x) : (y)) |
| Computes the maximum of x and y. | |
| #define | MIN(x, y) ((x) < (y) ? (x) : (y)) |
| Computes the minimum of x and y. | |
| #define | INDEX(i, j) ((i)*(nr+1)+(j)) |
| Computes the 1D index for concentration or alpha arrays, given pseudo-2D indices i (z index) and j (r index); uses column-major ordering. | |
Functions | |
| void | convolve3 (int M, int N, double *a, double scale1, double scale2, double *invr, double *out) |
Calculates updates to the concentration in a layer by applying the Laplacian in cylindrical coordinates; also scales by 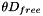 . More... . More... | |
| void | error (char *errorstring,...) |
| Print an error message to stderr and exit with status EXIT_FAILURE. More... | |
| void | print_usage_fit_layer (char *program) |
| Print usage message and exit with status EXIT_FAILURE. More... | |
| void | check_filename (char *in, char *out) |
| Make sure that the filename is not too long. More... | |
| double * | create_array (int N, char *string) |
| Create an array of doubles of the specified size. More... | |
| int | assemble_command (int argc, char *argv[], char *command) |
| Create a string containing the command used to run the program. More... | |
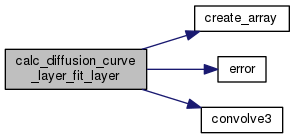
| void | calc_diffusion_curve_layer_fit_layer (int nt, int nz, int nr, int iprobe, int jprobe, int iz1, int iz2, int nolayer, double dt, double dr, double sd, double st, double alpha_so, double theta_so, double kappa_so, double alpha_sp, double theta_sp, double kappa_sp, double alpha_sr, double theta_sr, double kappa_sr, double dfree, double *t, double *s, double *invr, double *p) |
| Calculates the concentration as a function of space and time and returns the probe concentration as a function of time. More... | |
Header file for program fit-layer.
Definition in file header.h.
| int assemble_command | ( | int | argc, |
| char * | argv[], | ||
| char * | command | ||
| ) |
Create a string containing the command used to run the program.
It can be very useful for the program's output file to have a comment line that contains the command used to run the program. This function assembles that command into a string from the argc and argv parameters so it can be added to the output file.
| [in] | argc | Argument count (parameter passed to main) |
| [in] | argv | Argument vector (parameter passed to main) |
| [out] | command | String containing the command line |
| void calc_diffusion_curve_layer_fit_layer | ( | int | nt, |
| int | nz, | ||
| int | nr, | ||
| int | iprobe, | ||
| int | jprobe, | ||
| int | iz1, | ||
| int | iz2, | ||
| int | nolayer, | ||
| double | dt, | ||
| double | dr, | ||
| double | sd, | ||
| double | st, | ||
| double | alpha_so, | ||
| double | theta_so, | ||
| double | kappa_so, | ||
| double | alpha_sp, | ||
| double | theta_sp, | ||
| double | kappa_sp, | ||
| double | alpha_sr, | ||
| double | theta_sr, | ||
| double | kappa_sr, | ||
| double | dfree, | ||
| double * | t, | ||
| double * | s, | ||
| double * | invr, | ||
| double * | p | ||
| ) |
Calculates the concentration as a function of space and time and returns the probe concentration as a function of time.
This function solves the diffusion equation in each layer 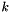 ,
,
![\[ \frac{\partial c_k(\vec r, t)}{\partial t} = D_{free} \theta_k \nabla^2{c_k(\vec r, t)} + s/\alpha_k - \kappa_k c_k(\vec r, t) \quad , \]](form_37.png)
subject to the continuity conditions at the interfaces between layers
![\[ c(\vec r_k, t) = c(\vec r_{k+1}, t) \quad \]](form_38.png)
![\[ \alpha_k \theta_k \nabla c(\vec r_k, t) = \alpha_{k+1} \theta_{k+1} \nabla c(\vec r_{k+1}, t) \quad \]](form_39.png)
and the boundary condition 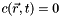 (total absorption) at the top, the bottom, and the side of the cylinder, where
(total absorption) at the top, the bottom, and the side of the cylinder, where
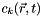 = concentration in layer
= concentration in layer 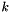
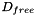 = free diffusion coefficient
= free diffusion coefficient
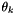 = permeability in layer
= permeability in layer 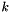
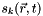 = source in layer
= source in layer 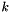
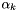 = extracellular volume fraction in layer
= extracellular volume fraction in layer 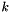
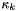 = nonspecific clearance factor in layer
= nonspecific clearance factor in layer 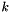
This function calls convolve3() to compute the Laplacian in cylindrical coordinates.
| [in] | nt | Number of support points in time |
| [in] | nz | Number of rows of concentration matrix |
| [in] | nr | Number of columns of concentration matrix |
| [in] | iprobe | z-index of probe location |
| [in] | jprobe | r-index of probe location |
| [in] | iz1 | z-index of SR-SP boundary |
| [in] | iz2 | z-index of SP-SO boundary |
| [in] | nolayer | Flag for whether to model a single homogeneous environment (true) or to model a 3-layer environment |
| [in] | dt | Spacing in time ( 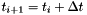 ) ) |
| [in] | dr | Spacing in r (in this program, 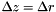 ) ) |
| [in] | sd | Source delay (time before source starts) |
| [in] | st | Duration of source |
| [in] | alpha_so | Extracellular volume fraction in SO layer |
| [in] | theta_so | Permeability in SO layer |
| [in] | kappa_so | Nonspecific clearance factor in SO layer |
| [in] | alpha_sp | Extracellular volume fraction in SP layer |
| [in] | theta_sp | Permeability in SP layer |
| [in] | kappa_sp | Nonspecific clearance factor in SP layer |
| [in] | alpha_sr | Extracellular volume fraction in SR layer |
| [in] | theta_sr | Permeability in SR layer |
| [in] | kappa_sr | Nonspecific clearance factor in SR layer |
| [in] | dfree | Free diffusion coefficient |
| [in] | t | Time array |
| [in] | s | Source array |
| [in] | invr | Array for 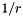 values values |
| [out] | p | Probe array (concentration as a function of time) |
Definition at line 94 of file model.c.
| void check_filename | ( | char * | in, |
| char * | out | ||
| ) |
Make sure that the filename is not too long.
If the input string is not to big, copy it to the output string. Otherwise, exit with an error message. This function is used to check that the user input filename is not too long.
| [in] | in | Input string = the user input filename (or base filename) |
| [out] | out | Output string = the input string, if not too long |
Definition at line 103 of file extras.c.
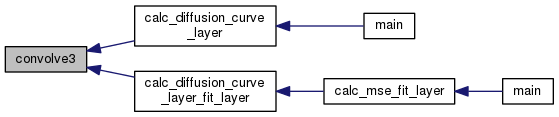
| void convolve3 | ( | int | M, |
| int | N, | ||
| double * | a, | ||
| double | scale1, | ||
| double | scale2, | ||
| double * | invr, | ||
| double * | out | ||
| ) |
Calculates updates to the concentration in a layer by applying the Laplacian in cylindrical coordinates; also scales by 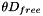 .
.
Calculates the update terms
![\[ \mathbf{out} = s_1 \mathbf{L} * \mathbf{a} + \frac{s_2}{r} \mathbf{D} * \mathbf{a} \qquad (r \neq 0) \]](form_25.png)
or
![\[ \mathbf{out} = s_1 \mathbf{L0} * \mathbf{a} \qquad (r = 0) \]](form_26.png)
where
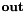 = output array
= output array
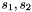 = scaling factors
= scaling factors
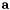 = input array (concentration)
= input array (concentration)
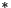 means convolution
means convolution
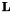 = 3x3 Laplace operator (see above)
= 3x3 Laplace operator (see above)
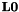 = 3x3 Laplace operator modified for
= 3x3 Laplace operator modified for 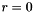
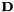 = 3x3 derivative operator for
= 3x3 derivative operator for 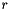
Note that the index 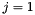 corresponds to
corresponds to 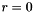 .
.
| [in] | M | Number of columns of input matrix (z) |
| [in] | N | Number of rows of input matrix (r) |
| [in] | a | Input matrix (concentration, c) |
| [in] | scale1 | Scaling factor #1 |
| [in] | scale2 | Scaling factor #2 |
| [in] | invr | Vector of 1/r values |
| [out] | out | Output matrix |
Definition at line 133 of file convo.c.
| double* create_array | ( | int | N, |
| char * | string | ||
| ) |
Create an array of doubles of the specified size.
Also initialize the array to all 0.
| [in] | N | Number of elements in array |
| [in] | string | String containing a description of the array (for debugging purposes) |
Definition at line 105 of file extras.c.
| void error | ( | char * | errorstring, |
| ... | |||
| ) |
 1.8.11
1.8.11